Deep learning object detection
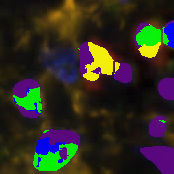
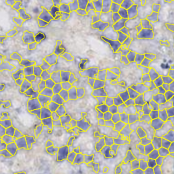
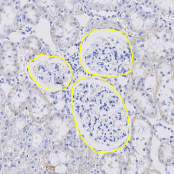
Version 3.0 integrates the deep learning framework Tensorflow for object detection. You can use a pre-trained model, like our glomeruli detection model available in our model zoo, or train your own deep learning model based on manually created object annotations. This new feature allows to detect arbitrary complex and heterogeneous objects like glomeruli, vessels, and[…]