Features
Sophisticated Image Analysis Algorithms
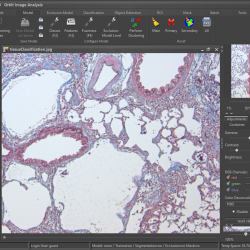
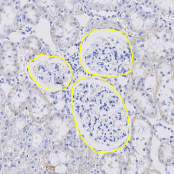
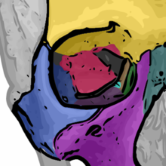
Orbit has many build-in image analysis algorithms. Tissue quantification using machine learning techniques, object / cell segmentation, and object classification are the basic ones. Region of interest (ROI) can be defined by manual annotations or via a trainable exclusion map. Everything can be combined.
WORKS FOR BIG IMAGES, e.g. WHOLE SLIDE SCANS
All algorithms are build to work on really big images, up to gigapixel images, especially whole slide scans.
This is possible due to a tile-based processing pipeline and combination with the use of different resolutions of the image.
Connectivity: OMERO & Spark connectors
You can use your existing image server, e.g. it has been designed to work great with Omero.
Orbit can also run in stand-alone mode and open whole slide images like SVS, NDPI, SCN, ….
A Spark infrastructure can be used as scaleout infrastructure to distribute computation intensive tasks.
Scripting & Extentions & Connectors
Orbit provides a versatile API for developers to create scripts or extentions. For instance, you can easily iterate over all tiles within a defined ROI and apply any algorithm or ImageJ plugin. You write s.th. which works for small in-memory images, Orbit takes care that is works for really big images.
Individual image server and scaleout connectors can also be created by implementing clear interfaces.
What it is
And it will stay for free.
No demo, it's a full version.
Orbit Image Analysis is a free open source software with the focus to quantify big images like whole slide scans.
It can connect to image servers, e.g. Omero.
Analysis can be done on your local computer or via scaleout functionality in a distrubuted computing environment like a Spark cluster.
Sophisticated image analysis algorithms incl. tissue quantification using machine learning, object segmentation and classification are build in. In addition a versatile API allows you to enhance Orbit and to run your own scripts.
-
Sophisticated Algorithms
-
Connectivity
-
Scaleout
-
Programming API
Image Analysis

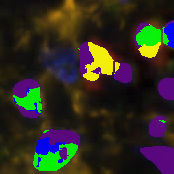
Tissue Quantification
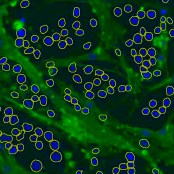
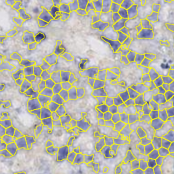
Object segmentation
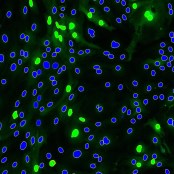
Object Classification